BUPID-PALM
BUPID-PALM: Boston University Protein Identifier – Parallel Acquisition Library Matcher
Ion mobility mass spectrometry offers separation capabilities complementary to LC on a time scale compatible with LC-MS; however, computational analysis of these data can be challenging. The addition of a second time dimension is problematic for many conventional LC-MS data analysis tools. The potential for co-isolation at the MS1 and MS2 level makes solving IM all-ion fragmentation (AIF) mass spectra difficult. Additionally, the need to identify glycopeptides introduces another layer of complexity in the analysis.
Building upon our experience with analyzing glycopeptide tandem mass spectra (GlycReSoft) and top-down data analysis (BUPID Top Down) we developed BUPID-PALM (Parallel Acquisition Library Matcher) for the analysis of data-independent analysis (DIA) all-ion fragmentation data (AIF) and MS2 data obtained from ion mobility mass spectrometry experiments.
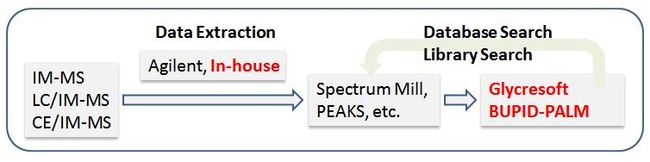
BUPID-PALM leverages the results of novel glycopeptide data analysis software tools and generates spectral libraries which can be used to interpret IM-AIF MS data. We first perform an LC-MS/MS run with a DDA method to create a series of reference spectra or spectral libraries and then re-analyze the sample under the same conditions with an IM-AIF MS method. The DDA data, which are computationally easier to process with existing software, serve as a spectral library against which we can match the DIA data. Since our goal is the assignment of glycopeptides, the DDA data are analyzed using GlycReSoft, a software package developed in our Center for Biomedical Mass Spectrometry at Boston University. Other search engines can be used to supplement or replace GlycReSoft, depending on the purpose of the analysis. An R script is used to translate the database search results into a simplified format that can be used by the IM-MS matching software.
If you are interested in evaluating BUPID-PALM, please contact Prof. Mark McComb.