GMS Research Symposium Award Winners
Congratulations to Nadia Mirza-Romero from the Harris lab & Diana Yeritsyan from the Layne lab who were graduate student winners at the 2026 GMS Research Symposium. Nadia's poster "Identifying PrPSc infection susceptibility and trafficking using regional in vitro stellate astrocyte cultures" was a winner in late career PhD or MD/PHD category. Diana received the research talk award in the early career category for her project "Functional characterization of ACLP mutations that cause Ehlers Danlos Syndrome". Congratulations Nadia (left) and Diana (right) receiving award from GMS Associate Dean Jamie McKnight!
Congratulations Cheyanne Frosti—GMS Outstanding Student Achievement Award
Congratulations to Cheyanne Frosti as a recipient of a GMS Outstanding Student Achievement Award. Cheyanne was awarded the Community Service Award in the PhD or MD/PhD category. This award recognizes graduating students whose contributions set them apart from their peers and Cheyanne will receive $500 and will be highlighted in the commencement programs. Congratulations Cheyanne!
Achievement Award. Cheyanne was awarded the Community Service Award in the PhD or MD/PhD category. This award recognizes graduating students whose contributions set them apart from their peers and Cheyanne will receive $500 and will be highlighted in the commencement programs. Congratulations Cheyanne!
New Publication from the Layne Lab: How the Extracellular Matrix Can Push Cells Toward Disease
Fibrosis is a major driver of organ dysfunction, characterized by excessive collagen accumulation and progressive tissue stiffening, yet effective treatment options remain limited. Understanding how changes in the extracellular matrix (ECM) influence cell behavior is key to identifying new therapeutic strategies.
In a new study from the Layne laboratory, Cheyanne Frosti and co-author Diana Yeritsyan identify a matrix-dependent mechanism by which aortic carboxypeptidase-like protein (ACLP) promotes fibrogenic activation. When bound to collagen, ACLP enhances β1 integrin activation and focal adhesion maturation, leading to downstream RhoA and Rac1 signaling, actin cytoskeletal remodeling, and nuclear accumulation of myocardin-related transcription factor A (MRTFA). These events drive transcriptional programs associated with ECM remodeling and cytoskeletal organization, promoting a pro-fibrotic state.
Importantly, this pathway highlights ACLP as a matrix-derived cue that links ECM composition to integrin-mediated mechanical signaling, providing insight into how physical changes in the ECM contribute to fibrotic disease progression and revealing new potential avenues for therapeutic targets.
Congratulations to Thomas Liontis
Congratulations to Thomas Liontis from the Grishok lab who successfully defended his PhD dissertation: Regulation of RNA Interference Pathways by Disruptor of Telomeric Silencing (DOT1) and Insulin/Insulin-Like Growth Factor-1 Signaling"
Congratulations to Jean Gatdula!
Congratulations to Jean Gatdula in the Harris lab for successfully defending his PhD dissertion; "MECHANISMS OF PRION SYNAPTOTOXICITY AND DEVELOPMENT OF A PRION PROTECTIVE VARIANT AS A THERAPEUTIC"
New publication from the Harris lab: Triggers for Prion toxicity
Prion diseases are fatal neurodegenerative disorders that affect both humans and animals. These diseases are caused by PrPSc, a misfolded and infectious isoform of the normal cellular prion protein (PrPC), which propagates by a self-templating mechanism. While considerable progress has been made in understanding prion propagation and infectivity, the early cellular events that initiate prion-induced synaptic loss and neurodegeneration remain poorly defined.
 A recent publication in PLoS Pathogens from the Harris lab, led by graduate student Jean Gatdula, focused on the initial molecular events on the neuronal surface that initiate prion synaptotoxicity. Using a specialized neuronal culture system that allows direct measurements of synaptic integrity, they found that experimental manipulations that blocked formation of pathogenic PrPSc on the neuronal surface prevented synaptic damage. Thus, membrane-attached PrPSc molecules are directly responsible for triggering a prion synaptotoxic signaling cascade, presumably by interacting with other proteins or lipids on the membrane surface. These results not only illuminate basic pathogenic mechanisms, but they suggest a novel therapeutic approach using PrP molecules that are locked in the PrPC conformation.
A recent publication in PLoS Pathogens from the Harris lab, led by graduate student Jean Gatdula, focused on the initial molecular events on the neuronal surface that initiate prion synaptotoxicity. Using a specialized neuronal culture system that allows direct measurements of synaptic integrity, they found that experimental manipulations that blocked formation of pathogenic PrPSc on the neuronal surface prevented synaptic damage. Thus, membrane-attached PrPSc molecules are directly responsible for triggering a prion synaptotoxic signaling cascade, presumably by interacting with other proteins or lipids on the membrane surface. These results not only illuminate basic pathogenic mechanisms, but they suggest a novel therapeutic approach using PrP molecules that are locked in the PrPC conformation.
Congratulations Cheyanne Frosti
Congratulations to Cheyanne Frosti in the Layne lab for successfully defending her PhD dissertion; "MECHANOREGULATION OF FIBROBLAST ACTIVATION BY AORTIC CARBOXYPEPTIDASE-LIKE PROTEIN"
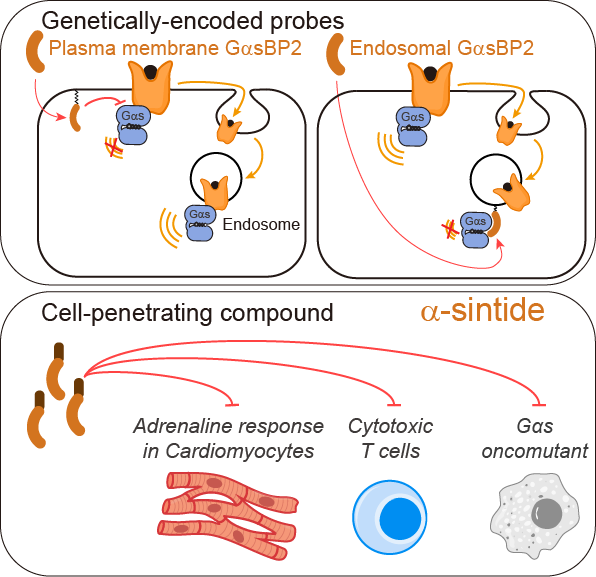
New publication from the Garcia-Marcos Lab: Inhibitory probes for spatiotemporal analysis of Gαs protein signaling
NEW PUBLICATION BY THE GARCIA-MARCOS LAB
A recent publication in Nature Chemical Biology (https://www.nature.com/articles/s41589-025-02138-1) led by Jingyi Zhao in the Garcia-Marcos Lab describes the discovery and optimization of broadly applicable genetically-encoded probes and peptide-based compounds specifically inhibit Gαs, the prototypical signal transducer of G protein-coupled receptors (GPCRs). These tools were leveraged to provide new mechanistic insights into GPCR signaling at the subcellular scale by revealing definitive evidence for G protein signaling at endosomes. The newly developed peptide-based compound named α-sintide allowed the inhibition signaling in multiple contexts: blocking an oncogenic G protein mutant, inhibiting heart cell responses to adrenaline, or preventing T cell exhaustion.
This work was done in collaboration with the Varelas Lab in our Department and with the Irannejad Lab and Vilardaga Lab at UCSF and U of Pittsburgh, respectively.
Congratulations to our BU Pilot Grant Awardees!
Congratulations to our BU Pilot Grant Awardees!
Please join us in congratulating the five Biochemistry & Cell Biology PIs who were selected for BU Pilot Grants:
-
Daniel Cifuentes – Spivack Neuroscience Pilot Grant
-
Valentina Perissi – Dahod Breast Cancer Research Pilot Grant
-
Nelson Lau – Sexual Medicine Pilot Grant
-
Bob Varelas – Dahod Breast Cancer Research Pilot Grant
-
Mike Blower – Shipley Prostate Cancer Pilot Grant
Well deserved—congratulations to all!
New research from the Cifuentes Lab: RBPscan, a quantitative in vivo tool for profiling RNA-binding protein interactions.
New research from the Cifuentes Lab: RBPscan, a quantitative in vivo tool for profiling RNA-binding protein interactions.
Researchers in the Cifuentes lab at Boston University have published a new study in Molecular Cell describing RBPscan, a method that enables quantitative measurement of protein-RNA interactions directly in living cells.
RNA-binding proteins regulate nearly every step of gene expression, but existing methods have made it difficult to determine where these proteins bind and how strongly they interact with RNA in their native cellular context. RBPscan was developed to address this challenge by combining RNA editing with massively parallel reporter assays to provide a simple, scalable, and quantitative readout of protein-RNA interactions in vivo.

The study demonstrates that RBPscan can identify both linear and structured RNA-binding motifs, quantify relative binding affinities across multiple experimental systems including human cells, zebrafish embryos, and yeast, and link binding strength to functional outcomes such as mRNA decay. The method also enables precise mapping of binding sites within full-length transcripts, providing positional information that complements existing approaches.
By making quantitative analysis of protein-RNA interactions more accessible, RBPscan provides a versatile new tool for studying post-transcriptional gene regulation and opens the door to discovering how RNA-binding proteins function in their native biological contexts.
Congratulations to the exceptional and dedicated Cifuentes lab team who made this work possible, with Dmitry Kretov spearheading the experimental work and now leading his own lab at CHU de Québec-Université Laval Research Center.
Publisher online February 6th, 2026.
Links to the paper:
https://authors.elsevier.com/a/1mZlb3vVUPZNXj
https://www.cell.com/molecular-cell/fulltext/S1097-2765(26)00023-7